A
new era in biology
opens
thanks to the
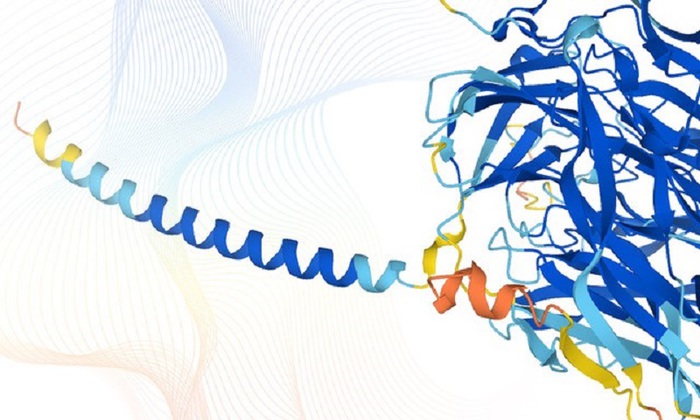
more complete and accurate 'portrait' of proteins
in the human body: this was created by the
AlphaFold
artificial intelligence system
, developed by the British company
DeepMind
from
, which managed to predict the 3D structure of
20,000 proteins
expressed by the human genome, in addition to those of 20 other organisms (from the mouse to the malaria parasite) crucial for research. The results, published in the journal Nature, are collected in a
database
that in collaboration with the European Molecular Biology Laboratory (EMBL) is made freely accessible to scientists from all over the world, to
accelerate research
in the life sciences, to
develop new drugs
and
make the most of the planet's resources
.
The database greatly increases the knowledge gained so far, more than
doubling
the number of human protein structures predicted with great accuracy. Previously, it was possible to determine the exact location of
17%
of the amino acids that make up human proteins: this result was achieved with decades of particularly long and complex laboratory studies and experiments. Artificial intelligence, on the other hand, managed in a few months to predict the position of 58% of amino acids with a good margin of certainty:
35.7%
was localized with a very high degree of confidence, practically double what was obtained experimentally. .
"This will be one of the most important datasets from human genome mapping," says EMBL Deputy Director General and EMBL-EBI Director Ewan Birney. "Making AlphaFold predictions accessible to the international scientific community opens up many new avenues of research, from neglected diseases to new enzymes for biotechnology and much more. It is a great new scientific tool, which complements existing technologies, and will allow us to broaden the boundaries of our understanding of the world ".


/cloudfront-eu-central-1.images.arcpublishing.com/prisa/XMI5DEX3DJDINGEKDTN43BWB5E.jpg)




